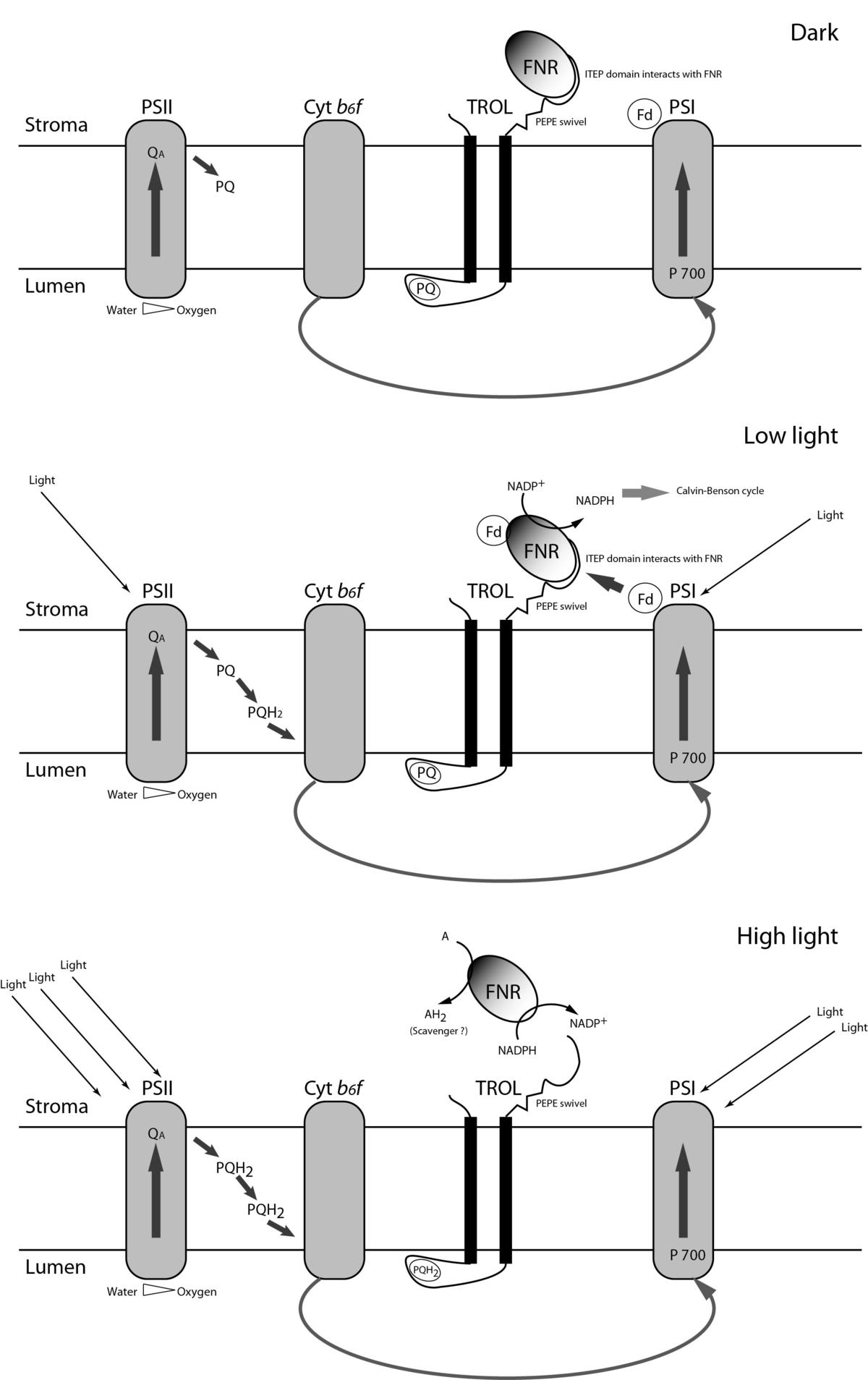
Mechanisms controlled by the TROL protein (Thylakoid Rhodanase-like)
We have recently cloned and characterized chloroplast protein TROL. TROL is an integral protein of thylakoid membranes located in the vicinity of the Photosystem I (PSI) and the cytochrome b6/f complex. TROL possesses several intriguing domains which perform different functions in the processes of electron transfer. At the center of the protein is the rhodanase-like domain (RHO). Rhodanases and the CDC25 phosphatases share common evolutionary origin. Rhodanase domains are involved in the metabolism of sulfur, while the CDC25 phosphatases regulate cell cycle in almost all eukaryotes.
At the carboxy terminus of the protein is the so called ITEP domain responsible for the binding of ferredoxin-NADP+ oxidoreductase, enzyme which catalyzes final electron transfer in photosynthesis, from ferredoxin to NADP+. RHO and ITEP are connected by the PEPE domain which is rich in proline. This domain probably serves a regulatory role in the dynamic binding of the FNR. We use knock-out plants for biochemical and biophysical analyses which allow us insight into complex processes of photosynthetic electron flow.
State-transitions regulated by the TLP40 protein
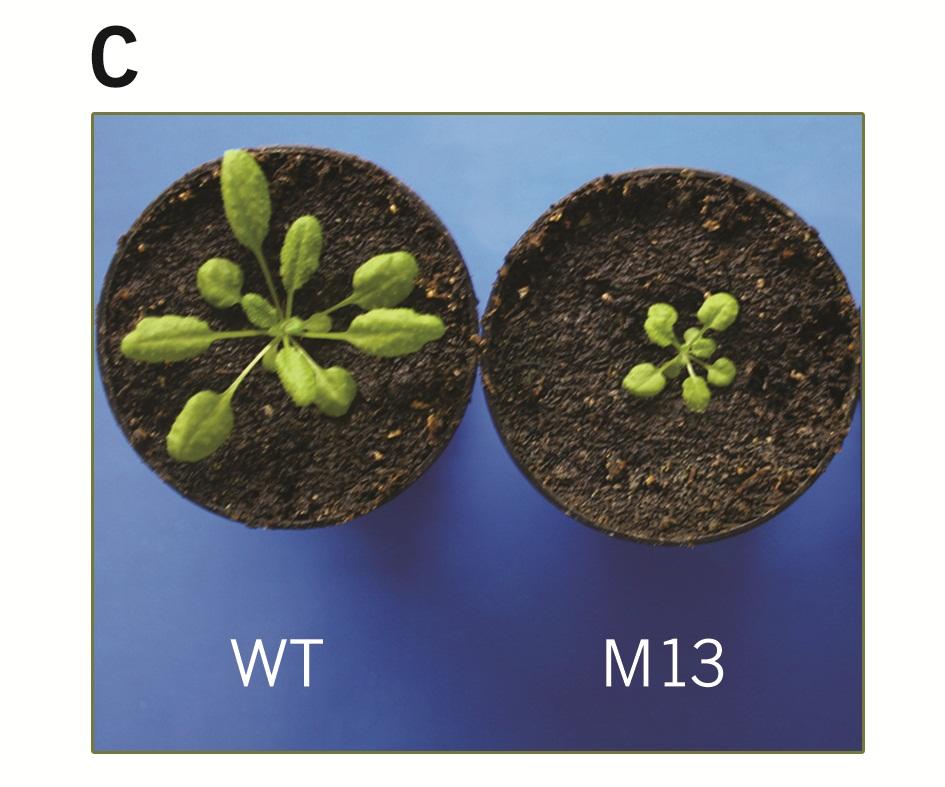
TLP40 is the first discovered complex immunophilin located in the thylakoid lumen. TLP40 and its orthologue CYP38 are involved in the regulation of protein dephosphorylation connected with the molecular mechanism of regulation known as the State-transitions. State transitions are regulated by the action of thylakoid protein kinases and phosphatases, as well as with the other auxiliary proteins. TLP40 forms dimers in the thylakiod lumen by utilizing the amino-terminal leucin-zipper domain. Arabidopsis thaliana plants which have knocked-out gene which encodes the TLP40 orthologue - CYP38 have stunned phenotype (M13 mutants).
Biophysical mesurements performed by using pulse amplified modulation (PAM) technique indicate that M13 plants are locked in the State II, i.g. dephosphorylation is impaired. Thus we have demonstrated that CYP38 controls the processes of dephosphorylation in vivo.
Metabolic engineering of cyanobacteria
Cyanobacteria Synechocistis PCC6803 are prokayotic ancestors of modern chloroplasts. Cyanobacteria have approximately 3500 genes and are thus much more suitable for investigations of the genes with unknown functions. Additionally, cyanobacteria perform processes of photosynthesis which are highly similar to those taking place in chloroplasts. However, cyanobacteria also utilize alternative mechanisms of energy production which are dependent on the activity of the enzymes hydrogenases. Hydrogen gas is a byproduct of the hydrogense activity. Development of the system for biological production of the hydrogen gas is at the center of our research on metabolic engineering of cyanobacteria.
Members of the Group:
Dr. sc. Lea Vojta, research associate
Dr. sc. Snježana Jurić, postdoc
Dr. sc. Ana Tomašić Paić, postdoc
Renata Hanzer, doctoral student
Dipl. inž. Lucija Horvat, t.a.
Marta Čorić, diploma student
Morana Novak, diploma student
Collaborations
Prof. dr. sc. Hrvoje Lepeduš, Agricultural Institute Osijek, dr. sc. Domagoj Šimić, Agricultural Institute Osijek, prof. dr. sc. Vera Cesar, Department of Biology, University "J.J. Strossmayera" in Osijeku, prof. dr. sc. Marko Jelić, "Marko Marulić" Polytechnics, Knin, doc. dr. sc. Željka Vidaković Cifrek, Department of Biology, PMF, Zagreb, prof. dr. sc. Mario Cifrek, FER, Zagreb
Prof. J.F. Allen, Queen Mary University of London, UK, Prof. Karin Krupinska, Christian-Albrechts Universitaet, Kiel, Njemačka, Prof. Bernd Zechmann, Institute of Plant Sciences, University of Graz, Austrija, PD Veronica Maurino, Henrich Heine Universitaet, Duesseldorf, Njemačka, Prof. Enrico Schleiff, Goethe Universitaet, Frankfurt, Njemačka