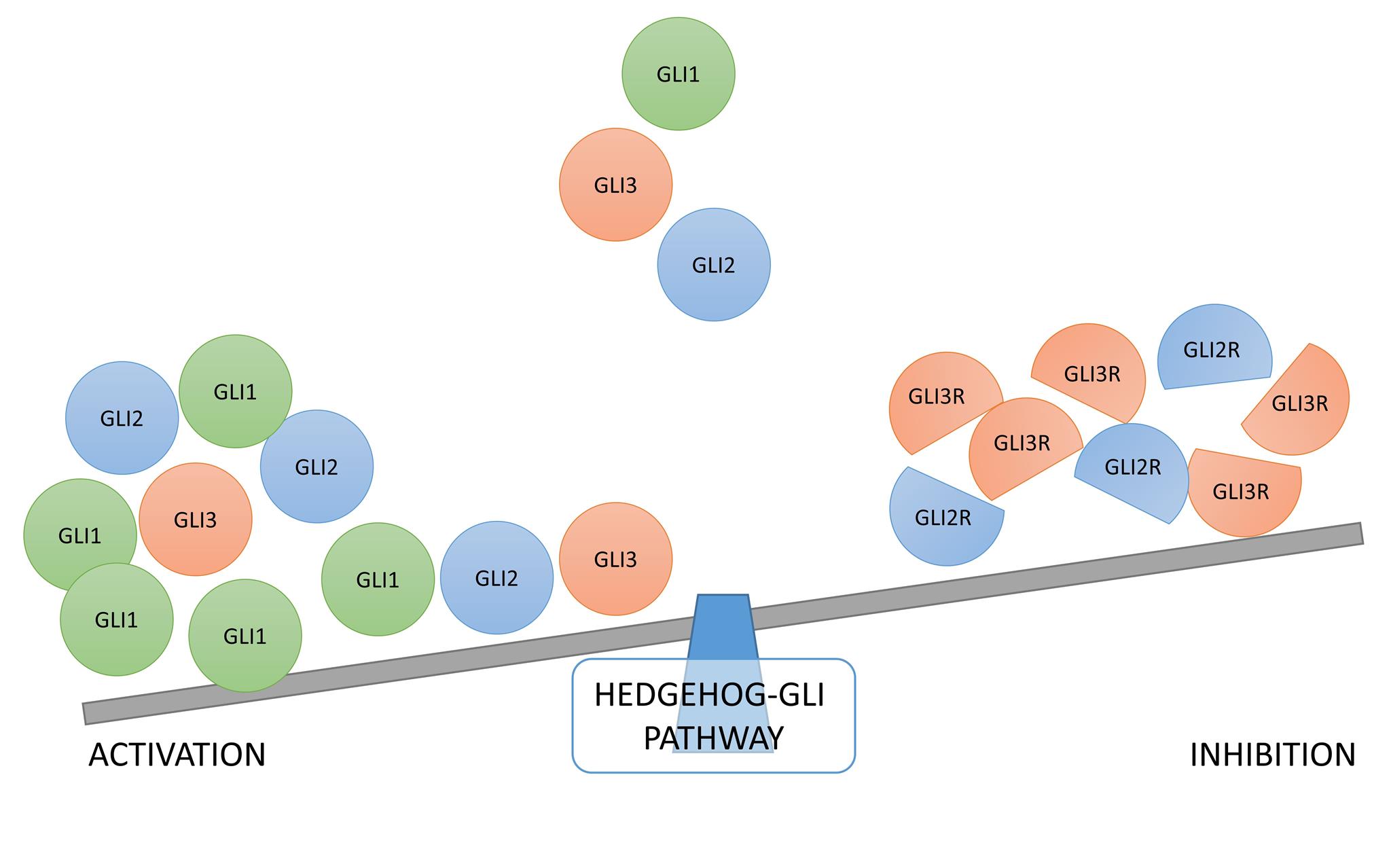
The Hedgehog signaling pathway has been implicated in development of various tumors, but the exact roles and molecular mechanisms are still not understood. The focus of this research are the tumors harboring BRAF or NRAS mutations, and their differential response to Hedgehog pathway inhibition. Both BRAF and NRAS non-canonically activate the three GLI proteins, but the activity and interplay of this interaction is still unclear. With this project we propose to determine the exact transcriptional targets of GLI1, GLI2 and GLI3 proteins in melanoma with different genetic backgrounds, either harboring a BRAF mutation, a NRAS mutation, or no mutation in these two genes. For this purpose, we will first perform ChIP-seq to determine the transcriptional targets, then knock-out each of the three GLI proteins and sequence the transcriptomes of the knock-out cell lines. After the knock-outs have been generated, a series of cell assays will be performed to determine the role of each of the three GLI proteins in BRAF/NRAS dependent tumors, and to elucidate the mechanism of their action in melanoma. Combined treatments with Hedgehog inhibitors and specific BRAF/NRAS inhibitors, followed by generation and testing of resistant cell lines, will demonstrate if there is a potential benefit in the clinical management of melanoma. Finally, in the final year of the project, a set of clinical samples will be analyzed for these potential biomarkers to determine their possible use in the clinic. In summary, this project plans a complete analysis of the GLI code in BRAF/NRAS dependent melanoma, from determining the transcriptional targets and finding molecular mechanisms, to potential translation to the clinic.
Journal articles
1. Piteša, Nikolina; Kurtović, Matea; Bartoniček, Nenad; Gkotsi, Danai S, Čonkaš, Josipa; Petrić, Tina; Musani, Vesna; Ozretić, Petar; Riobo-Del Galdo, Natalia A; Sabol, Maja. Signaling switching from Hedgehog-GLI to MAPK signaling potentially serves as a compensatory mechanism in melanoma cell lines resistant to GANT-61. Biomedicines (2023), 11, 1353.
2. Čonkaš, Josipa; Sabol, Maja; Ozretić, Petar. "Toxic masculinity": what is known about the role of androgen receptors in head and neck squamous cell carcinoma. International Journal of Molecular Sciences (2023), 24, 3766.
3. Kurtović, Matea; Piteša, Nikolina; Bartoniček, Nenad; Ozretić, Petar; Musani, Vesna; Čonkaš, Josipa; Petrić, Tina; King, Cecile; Sabol, Maja
RNA-seq and ChIP-seq Identification of Unique and Overlapping Targets of GLI Transcription Factors in Melanoma Cell Lines. Cancers, 14 (2022), 18; 4540. doi:10.3390/cancers14184540
4. Trnski, Diana; Sabol, Maja; Tomić, Sanja; Štefanac, Ivan; Mrčela, Milanka; Musani, Vesna; Rinčić, Nikolina; Kurtović, Matea; Petrić, Tina; Levanat, Sonja; Ozretić, Petar
SHH-N non-canonically sustains androgen receptor activity in androgen-independent prostate cancer cells. Scientific reports, 11 (2021), 1; 14880, 11. doi:10.1038/s41598-021-93971-6
5. Zubčić*, Vedran; Rinčić*, Nikolina; Kurtović, Matea; Trnski, Diana; Musani, Vesna; Ozretić, Petar; Levanat, Sonja; Leović, Dinko; Sabol, Maja
GANT61 and Lithium Chloride Inhibit the Growth of Head and Neck Cancer Cell Lines Through the Regulation of GLI3 Processing by GSK3beta. International journal of molecular sciences, 21 (2020), 6410, 13 doi:10.3390/ijms21176410
Conference abstracts
1. Križanac, Marinela ; Leović, Matko ; Rinčić, Nikolina ; Trnski, Diana ; Musani, Vesna ; Ozretić, Petar ; Levanat, Sonja ; Zubčić, Vedran ; Leović, Dinko ; Sabol, Maja. Hedgehog-GLI signaling controls proliferation and invasiveness of head and neck squamous cell carcinoma. HDBMB2019 Crossroads in Life Sciences, 2019. (poster)
2. Rinčić, Nikolina ; Bartoniček, Nenad; Kurtović, Matea; Musani, Vesna; Trnski, Diana; Ozretić, Petar; Sabol, Maja. GLI transcription targets in melanoma cell lines. 45th FEBS Virtual Congress “Molecules of Life: Towards New Horizons“, 2021. (poster)
3. Rinčić, Nikolina ; Bartoniček, Nenad ; Kurtović, Matea ; Musani, Vesna ; Trnski, Diana ; Ozretić, Petar ; Sabol, Maja. Specific and overlapping GLI transcription targets in melanoma. EACR 2021 Virtual Congress, 2021. (poster)
4. Kurtović, Matea ; Rinčić, Nikolina ; , Trnski, Diana ; Musani, Vesna ; Ozretić, Petar ; Sabol, Maja. Generation of GLI1, GLI2 and GLI3 knock-out melanoma cell lines via CRISPR/Cas9 gene editing. EACR 2021 Virtual Congress, 2021. (poster)
5. Kurtović, Matea ; Rinčić, Nikolina ; Trnski, Diana ; Musani, Vesna ; Ozretić, Petar ; Sabol, Maja. CRISPR/Cas9 mediated knockout of GLI1, GLI2 and GLI3 genes in melanoma cell lines. 20th FEBS Young Scientists´Forum, 2021. (poster)
6. Kurtović, Matea ; Rinčić, Nikolina ; Trnski, Diana ; Musani, Vesna ; Ozretić, Petar ; Sabol, Maja. CRISPR/Cas9 mediated knockout of GLI1, GLI2 and GLI3 genes in melanoma cell lines. 45th FEBS Congress, 2021. (poster)
7. Petrić, Tina ; Trnski, Diana ; Kurtović, Matea ; Rinčić, Nikolina ; Musani, Vesna ; Ozretić, Petar ; Sabol, Maja. Hedgehog-GLI signaling pathway in 3D model of prostate cancer. Goodbye Flat Biology: Next Generation Cancer Models, 2021. (poster)
8. Piteša, Nikolina ; Bartoniček, Nenad ; Kurtović, Matea ; Musani, Vesna ; Trnski, Diana ; Ozretić, Petar ; Sabol, Maja. Genes for competing endogenous RNAs as targets of transcription factors GLI in melanoma cell lines. 5th Congress of the Serbian Association for Cancer Research (SDIR), 2021. (lecture)
9. Čonkaš, Josipa ; Josić, Janja ; Piteša, Nikolina ; Kurtović, Matea ; Petrić, Tina ; Musani, Vesna ; Sabol, Maja ; Vugrinec, Ozren ; Leović, Dinko ; Ozretić, Petar. The role of androgen and androgen receptors in metastasis of head and neck tumors. 6. Simpozij studenata doktorskih studija PMF-a, 2022. (poster)
10. Piteša, N ; Kurtović, M ; Bartoniček, N ; Petrić, T ; Čonkaš, J ; Musani, V ; Ozretić, P ; Sabol, M. Hedgehog-GLI and MAPK signalling pathway activity in GANT61-resistant melanoma cells. 28th Congress of the European Association for Cancer Research (EACR 2022 Congress) "Innovative Cancer Science: Translating Biology to Medicine", 2022. (poster)
11. Kurtović, Matea ; Bartoniček, Nenad ; Piteša, Nikolina ; Ozretić, Petar ; Musani, Vesna ; Petrić, Tina ; Čonkaš, Josipa ; Sabol, Maja. RNA sequencing reveals unique and overlapping transcriptional targets of GLI1, GLI2 and GLI3 in melanoma cell lines. 28th Congress of the European Association for Cancer Research (EACR 2022 Congress) "Innovative Cancer Science: Translating Biology to Medicine", 2022. (poster)
12. Piteša, Nikolina ; Bartoniček, Nenad ; Kurtović, Matea ; Petrić, Tina ; Čonkaš, Josipa ; Musani, Vesna ; Ozretić, Petar ; Sabol, Maja. Primary cilia - a potential link between hedgehog- GLI and MAPK signaling in GANT61 -resistant melanoma cell lines. HDBMB22: From Science to Knowledge, 2022. (poster)
13. Čonkaš, Josipa ; Josić, Janja ; Piteša, Nikolina ; Kurtović, Matea ; Petrić, Tina ; Musani, Vesna ; Sabol, Maja ; Vugrinec, Ozren ; Leović, Dinko ; Ozretić, Petar. Membrane androgen receptor OXER1 indicates a potential role in metastasis of head and neck tumors. 6th Meeting of the Croatian Association for Cancer Research with International Participation: Targeting Cancer (HDIR-6), 2022. (poster)
14. Piteša, Nikolina ; Bartoniček, Nenad ; Kurtović, Matea ; Petrić, Tina ; Čonkaš, Josipa ; Musani, Vesna ; Ozretić, Petar ; Sabol, Maja. Signaling Switch from Hedgehog-GLI to MAPK Potentially Drives Primary Cilia Loss in NRAS Mutated GANT61-Resistant Melanoma Cell Line. 6th Meeting of the Croatian Association for Cancer Research with International Participation: Targeting Cancer (HDIR-6), 2022. (poster)
15. Petrić, Tina ; Kurtović, Matea ; Piteša, Nikolina ; Čonkaš, Josipa ; Musani, Vesna ; Ozretić, Petar ; Sabol, Maja. Hedgehog-GLI signaling pathway in 3D model of prostate cancer. HDBMB22: From Science to Knowledge, 2022. (poster)
16. Petrić, Tina ; Kurtović, Matea ; Piteša, Nikolina ; Čonkaš, Josipa ; Musani, Vesna ; Ozretić, Petar ; Sabol, Maja. 3D culture of prostate cancer as a model to study hedgehog-gli signaling pathway. 6th Meeting of the Croatian Association for Cancer Research with International Participation: Targeting Cancer (HDIR-6), 2022. (poster)
17. Petrić, Tina. Let's go 3D: the future of prostate cancer models. Digital science conference Kemomind, 2022. (lecture)
18. Kurtović, Matea ; Piteša, Nikolina ; Bartoniček, Nenad ; Ozretić, Petar ; Musani, Vesna ; Petrić, Tina ; Čonkaš, Josipa ; Sabol, Maja. Unique and overlapping transcriptional targets of GLI1, GLI2 and GLI3 in melanoma cell lines. 6th Meeting of the Croatian Association for Cancer Research with International Participation: Targeting Cancer (HDIR-6), 2022. (poster)
19. Piteša, Nikolina ; Bartoniček, Nenad ; Kurtović, Matea ; Petrić, Tina ; Čonkaš, Josipa ; Musani, Vesna ; Ozretić, Petar ; Sabol, Maja. Characterization of GANT61 Resistant Cell Lines reveals new insights of Hedgehog-GLI and RAS/RAF/MAPK interplay in melanoma. EACR-Boehringer Ingelheim Drugging and Regulating the MAP Kinase Pathway, 2023. (poster)
20. Čonkaš, Josipa ; Josić, Janja ; Kranjčević, Jacqueline-Katrin ; Piteša, Nikolina ; Kurtović, Matea ; Musani, Vesna ; Sabol, Maja ; Vugrinec, Ozren ; Mumlek, Ivan ; Kvolik Pavić, Ana ; Zubčić, Vedran ; Leović, Dinko ; Ozretić, Petar. Correlation between the sex hormone receptors expression and demographic parameters in HNSCC patients 7. Simpozij studenata doktorskih studija PMF-a, 2023. (poster)
21. Piteša, Nikolina ; Kurtović, Matea ; Bartoniček, Nenad ; Gkotsi, Danai S, Čonkaš, Josipa ; Petrić, Tina ; Musani, Vesna ; Ozretić, Petar ; Riobo-Del Galdo, Natalia A ; Sabol, Maja. First Insights into the Characterization of GANT61 - Resistant Melanoma Cell Lines. Hedgehog signaling: From molecular structure to developmental biology and diseases, 2023. (poster)
22. Kurtović, Matea ; Bartoniček, Nenad ; Piteša, Nikolina ; Ozretić, Petar ; Musani, Vesna ; Čonkaš, Josipa ; Sabol, Maja. Unique and overlapping transcriptional targets of GLI1, GLI2 and GLI3 in melanoma cell lines. Hedgehog signaling: From molecular structure to developmental biology and diseases, 2023. (poster)
23. Sabol, Maja. Signalni putevi u melanomu: matičnost, plastičnost i rezistencija. 8. simpozij Apoptoza i novotvorine, 2023. (invited lecture)
24. Piteša, Nikolina; Kurtović, Matea; Čonkaš, Josipa; Musani, Vesna; Ozretić, Petar; Sabol, Maja. The role of Hedgehog signaling pathway in plasticity, stemness and resistance of melanoma. 6th Congress of the Serbian Association for Cancer Research "SDIR-6" with international participation "From Collaboration to Innovation in Cancer Research", 2023. (invited lecture)
25. Čonkaš, Josipa; Josić, Janja; Kranjčević, Jacqueline-Katrin; Milutin Gašperov, Nina; Piteša, Nikolina; Kurtović, Matea; Musani, Vesna; Sabol, Maja; Vugrinec, Ozren; Mumlek, Ivan et al. Expression Profile of Sex Hormone Receptors in Head and Neck Cancer: Unraveling Gender Disparities. 6th Congress of the Serbian Association for Cancer Research "SDIR-6" with international participation "From Collaboration to Innovation in Cancer Research", 2023. (poster)
Theses
1. Kurtović, Matea (2023) „Unique and overlapping target genes of transcription factors GLI1, GLI2 and GLI3 in human melanoma“, University of Rijeka, Doctoral study of medicinal chemistry, PhD thesis
2. Piteša, Nikolina (2023) „Interaction of RAS/RAF/MAPK and Hedgehog-GLI signaling pathways in therapy resistance in human melanoma cell lines“, Josip Juraj Strossmayer University of Osijek, Interdisciplinary doctoral study of molecular biosciences, PhD thesis
3. Brtan, Lucija (2023) “Construction and validation of the expression vector for the repressor form of the GLI3 protein in human cells“,University of Zageb, Faculty of Science, masters thesis
4. Čonkaš, Josipa (2021) “Construction of vectors for SHH protein expression in CHL-1 human melanoma cell line“, University of Zagreb, Faculty of Science, masters thesis
5. Brtan, Ana (2021) “Characterization and validation of SHH knock-in melanoma cell lines“, University of Zagreb, Faculty of Science, masters thesis
Vučemilo, Nikolina (2021) “Establishing a three-dimensional spheroid culture of head and neck tumor cells“, University of Zagreb, Faculty of Science, masters thesis
6. Rinčić, Nikolina (2018) “CRISPR/Cas9-mediated generation of GLI1 knock out cell line“, University of Zagreb, Faculty of Science, masters thesis
Awards
1. Piteša, Nikolina; Kurtović, Matea; Trsnki, Diana; Musani, Vesna; Ozretić Petar; Levanat, Sonja; Sabol, Maja. Annual award of the Ruđer Bošković Institute for best papers published in 2021: „GANT61 and Lithium chloride inhibit the growth of head and neck cancer cell lines through the regulation of GLI3 processing by GSK3beta“ by Zubčić et al.
2. Čonkaš, Josipa. Zoran Zgaga Award, awarded by the Croatian Association of Genetic Engineers, for the best masters thesis in academic year 2020/2021, titled “Construction of vectors for SHH protein expression in CHL-1 human melanoma cell line“
GLIcode